SSC/FreeEnergy/PolytropesEmbedded/Pt3C
Free Energy of Embedded Polytropes[edit]
Part I: Synopsis |
Part II: Truncated Polytropes |
Part III: Free-Energy of Bipolytropes |
||
IIIA: Focus on (5, 1) Bipolytropes |
IIIB: Focus on (0, 0) Bipolytropes |
IIIC: Overview |
||
Free-Energy of Bipolytropes[edit]
In this case, the Gibbs-like free energy is given by the sum of four separate energies,
|
|
|
|
In addition to specifying (generally) separate polytropic indexes for the core, , and envelope, , and an envelope-to-core mean molecular weight ratio, , we will assume that the system is fully defined via specification of the following five physical parameters:
- Total mass, ;
- Total radius, ;
- Interface radius, , and associated dimensionless interface marker, ;
- Core mass, , and associated dimensionless mass fraction, ;
- Polytropic constant in the core, .
In general, the warped free-energy surface drapes across a five-dimensional parameter "plane" such that,
|
|
|
|
Overview[edit]
BiPolytrope51[edit]
Key Analytic Expressions[edit]
|
Out-of-Equilibrium, Free-Energy Expression for BiPolytropes with |
|||
|---|---|---|---|
|
where,
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
From the accompanying Table 1 parameter values, we also can write,
|
|
|
|
|
|
|
|
Consistent with our generic discussion of the stability of bipolytropes and the specific discussion of the stability of bipolytropes having , it can straightforwardly be shown that is satisfied by setting ; that is, the equilibrium condition is,
|
|
|
|
|
|
|
|
where the last expression has been cast into a form that more clearly highlights overlap with the expression, below, for the equilibrium radius for zero-zero bipolytropes. Furthermore, the equilibrium configuration is unstable whenever,
that is, it is unstable whenever,
|
|
|
|
Table 1 of an accompanying chapter — and the red-dashed curve in the figure adjacent to that table — identifies some key properties of the model that marks the transition from stable to unstable configurations along equilibrium sequences that have various values of the mean-molecular weight ratio, .
Behavior of Equilibrium Sequence[edit]
Here we reprint Figure 1 from an accompanying chapter wherein the structure of five-one bipolytropes has been derived. It displays detailed force-balance sequences in the plane for a variety of choices of the ratio of mean-molecular-weights, , as labeled.
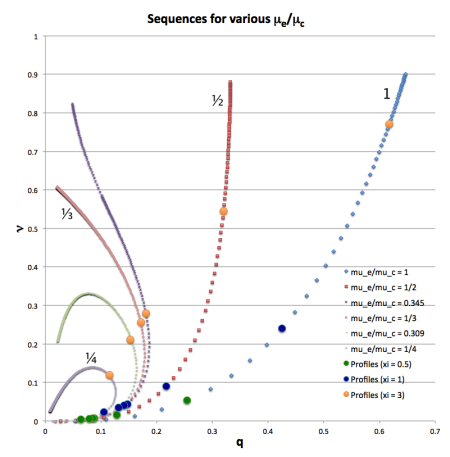
Limiting Values[edit]
Each sequence begins at the origin, that is, at . As , however, the sequences terminate at different coordinate locations, depending on the value of . In deriving the various limits, it will be useful to note that,
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Examining the three relevant parameter regimes, we see that:
- For , that is, …
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
and |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
- For , that is, …
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
and |
|
|
|
|
|
|
|
|
|
- For , that is, …
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
and |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Summarizing:
- For , that is, …
- For , that is, …
- For , that is, …
Turning Points[edit]
Let's identify the location of two turning points along the sequence — one defines and the other identifies . They occur, respectively, where,
|
|
and |
|
In deriving these expressions, we will use the relations,
|
|
|
|
|
|
|
|
where,
Given that,
|
|
|
|
we find,
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
And, given that,
|
|
|
|
we find,
|
|
|
|
|
|
|
|
In summary, then, the turning point occurs where,
|
|
|
|
and the turning point occurs where,
|
|
|
|
|
|
|
|
|
|
|
|
|
NOTE: As we show above, for the special case of — that is, when , precisely — the equilibrium sequence (as ) intersects the axis at precisely the value, . As is illustrated graphically in Figure 1 of an accompanying chapter, no turning point exists for values of . |
For the record, we repeat, as well, that the transition from stable to dynamically unstable configurations occurs along the sequence when,
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
In order to clarify what equilibrium sequences do not have any turning points, let's examine how the turning-point expression behaves as .
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
The leading-order term is unity on both sides of this expression, so they cancel; let's see what results from keeping terms .
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
We therefore conclude that the turning point does not appear along any sequence for which,
|
|
|
|
|
|
|
|
| Five-One Bipolytrope Equilibrium Sequences in Plane | |
|
Full Sequences for Various |
Magnified View with Turning Points and Stability Transition-Points Identified |
Graphical Depiction of Free-Energy Surface[edit]
| Figure 1: Free-Energy Surface for and | ||||
| ||||
For purposes of reproducibility, it is incumbent upon us to clarify how the values of the free energy were normalized in order to produce the free-energy surface displayed in Figure 1. The normalization steps are explicitly detailed within the fortran program, below, that generated the data for plotting purposes; here we provide a brief summary. We evaluated the normalized free energy, , across a zone grid of uniform spacing, covering the following domain:
|
|
|
|
|
|
|
|
(With this specific definition of the y-coordinate grid, is associated with zone 70.) After this evaluation, a constant, , was added to in order to ensure that the free energy was negative across the entire domain. Then (inorm = 1), for each specified interface location, , employing the equilibrium value of the free energy,
the free energy was normalized across all values of via the expression,
Finally, for plotting purposes, at each grid cell vertex ("vertex") — as well as at each grid cell center ("cell") — the value of the free energy, , was renormalized as follows,
Via this last step, the minimum "vertex" energy across the entire domain was 0.0 while the maximum "vertex" energy was 1.0.
| FORTRAN Program Documentation | Example Evaluations(See also associated Table 1) | ||||
|---|---|---|---|---|---|
| Coord. Axis | Coord. Name | Associated Physical Quantity | |||
| x-axis | bsize | ||||
| y-axis | csize | ||||
| Relevant Lines of Code | |||||
eta = 3.0d0*muratio*bsize/(1.0d0+bsize**2)
Gami = 1.0d0/eta-bsize
frakL = (bsize**4-1.0d0)/bsize**2 + &
& DATAN(bsize)*((1.0d0+bsize**2)/bsize)**3
frakK = Gami/eta + ((1.0d0+Gami**2)/eta)*(pii/2.0d0+DATAN(Gami))
E0 = ((5.0d0*frakL) + (4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK))/bsize**2+Efudge
fescalar(j,k) = (csize**(-0.6d0)*(5.0d0*frakL)&
& + csize**(-3.0d0)*(4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK)/csize)/bsize**2 + Efudge
if(inorm.eq.1)fescalar(j,k)=fescalar(j,k)/DABS(E0) &
& - E0/DABS(E0)
|
|||||
| Variable | Represents | Value calculated via the expression … | |||
| eta |
|
||||
| Gami | |||||
| frakL | |||||
| frakK | |||||
| E0 - Efudge |
|
||||
| Figure 2: Free-Energy Surface for and | ||
|
BiPolytrope00[edit]
|
Out-of-Equilibrium, Free-Energy Expression for BiPolytropes with Structural |
|||
|---|---|---|---|
|
where,
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
The associated equilibrium radius is,
|
|
|
|
We have deduced that the system is unstable if,
|
|
|
|
Fortran Code[edit]
This is the program that generated the free-energy data for the "five-one" bipolytrope that is displayed in the above, Figure 1 image/animation.
PROGRAM BiPolytrope
real*8 pii
real*8 bmin,bmax,cmin,cmax,db,dc
real*8 c(200),b(200),chalf(199),bhalf(199)
real*8 bsize,csize,emin,emax
real*8 fepoint(200,200),fescalar(199,199)
real*8 ell(200),ellhalf(199)
real*8 muratio,eta,Gami,frakK,frakL
real*8 eshift,ediff
real xx(200),yy(200),cell(199,199),vertex(200,200)
real valuemin,minlog,valufudge
real*8 q,nu,chiEq,Enorm,E0,Efudge
integer j,k,n,nmax,inorm
101 format(4x,'bsize',7x,'csize',8x,'xi',10x,'A',12x,'B',12x,&
&'fM',13x,'fW',11x,'fA',/)
! 102 format(1p8d12.4)
103 format(2i5,1p3d14.6)
201 format(5x,'valuemin = ',1pe15.5,//)
205 format(5x,'For Cell-center ... emin, emax = ',1p2d14.6,/)
206 format(5x,'For Cell-vertex ... emin, emax = ',1p2d14.6,/)
701 format(5x,1p10d10.2)
700 format(8x,'xi',9x,'ell',8x,'eta',8x,'Lambda',5x,'frakK',&
& 5x,'frakL',8x,'q',5x,'nu',5x,'chiEq',8x,'E0',/)
!!!!!!!!
!!!!!!!!
inorm=1
pii = 4.0d0*datan(1.0d0)
muratio = 1.0d0
bsize = 0.0d0
csize = 0.0d0
Efudge = -10.0d0
write(*,101)
! write(*,102)bsize,csize,xival,coefA,coefB,fM,fW,fA
!!!!!!!!!!!
!
! In this free-energy routine, c = X = chi/chi_eq and b = xi_i
!
!!!!!!!!!!!
nmax = 200
bmin = 1.0d0
bmax = 3.0d0
db = (bmax-bmin)/dfloat(nmax-1)
b(1) = bmin
ell(1) = b(1)/dsqrt(3.0d0)
! These values of cmin and cmax ensure that X=1 occurs at zone 70
cmin = 0.469230769d0
cmax = 2.00d0
dc = (cmax-cmin)/dfloat(nmax-1)
c(1) = cmin
do n=2,nmax
b(n) = b(n-1)+db
c(n) = c(n-1)+dc
ell(n) = b(n)/dsqrt(3.0d0)
enddo
do n=1,nmax-1
bhalf(n) = 0.5d0*(b(n)+b(n+1))
chalf(n) = 0.5d0*(c(n)+c(n+1))
ellhalf(n) = bhalf(n)/dsqrt(3.0d0)
enddo
!
! BEGIN LOOP to evaluate free energy (cell center)
!
emin = 0.0d0
emax = 0.0d0
write(*,700)
do j=1,nmax-1
bsize = ellhalf(j)
eta = 3.0d0*muratio*bsize/(1.0d0+bsize**2)
Gami = 1.0d0/eta-bsize
frakL = (bsize**4-1.0d0)/bsize**2 + &
& DATAN(bsize)*((1.0d0+bsize**2)/bsize)**3
frakK = Gami/eta + ((1.0d0+Gami**2)/eta)*(pii/2.0d0+DATAN(Gami))
q = 1.0d0/(1.0d0 + (0.5d0*pii+DATAN(Gami))/eta)
nu = bsize*q/dsqrt(1.0d0+Gami**2)
chiEq = dsqrt(pii/8.0d0)*(nu**2/(q*bsize**2))&
& *((1.0d0+bsize**2)/(3.0d0*bsize))**3
Enorm = 16.0d0*(q/nu**2)*chiEq
E0 = ((5.0d0*frakL) + (4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK))/bsize**2+Efudge
write(*,701)b(j),bsize,eta,Gami,frakK,frakL,q,nu,chiEq,E0
do k=1,nmax-1
csize=chalf(k)
fescalar(j,k) = (csize**(-0.6d0)*(5.0d0*frakL)&
& + csize**(-3.0d0)*(4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK)/csize)/bsize**2 + Efudge
if(inorm.eq.1)fescalar(j,k)=fescalar(j,k)/DABS(E0) &
& - E0/DABS(E0)
if(fescalar(j,k).gt.0.5d0)fescalar(j,k)=0.5d0
if(fescalar(j,k).lt.emin)emin=fescalar(j,k)
if(fescalar(j,k).gt.emax)emax=fescalar(j,k)
! write(*,103)j,k,bsize,csize,fescalar(j,k)
enddo
enddo
write(*,205)emin,emax
!
! BEGIN LOOP to evaluate free energy (cell vertex)
!
emin = 0.0d0
emax = 0.0d0
do j=1,nmax
bsize = ell(j)
eta = 3.0d0*muratio*bsize/(1.0d0+bsize**2)
Gami = 1.0d0/eta-bsize
frakL = (bsize**4-1.0d0)/bsize**2 + &
& DATAN(bsize)*((1.0d0+bsize**2)/bsize)**3
frakK = Gami/eta + ((1.0d0+Gami**2)/eta)*(pii/2.0d0+DATAN(Gami))
q = 1.0d0/(1.0d0 + (0.5d0*pii+DATAN(Gami))/eta)
nu = bsize*q/dsqrt(1.0d0+Gami**2)
chiEq = dsqrt(pii/8.0d0)*(nu**2/(q*bsize**2))&
& *((1.0d0+bsize**2)/(3.0d0*bsize))**3
Enorm = 16.0d0*(q/nu**2)*chiEq
E0 = ((5.0d0*frakL) + (4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK))/bsize**2 + Efudge
do k=1,nmax
csize=c(k)
fepoint(j,k) = (csize**(-0.6d0)*(5.0d0*frakL)&
& + csize**(-3.0d0)*(4.0d0*frakK)&
& - (3.0d0*frakL+12.0d0*frakK)/csize)/bsize**2 + Efudge
if(inorm.eq.1)fepoint(j,k)=fepoint(j,k)/DABS(E0) &
& - E0/DABS(E0)
if(fepoint(j,k).gt.0.5d0)fepoint(j,k)=0.5d0
if(fepoint(j,k).lt.emin)emin=fepoint(j,k)
if(fepoint(j,k).gt.emax)emax=fepoint(j,k)
! write(*,103)j,k,bsize,csize,fepoint(j,k)
enddo
enddo
write(*,206)emin,emax
!
! Now fill single-precision arrays for plotting.
!
do n=1,nmax
! xx(n)=b(n)/b(nmax)
! yy(n)=c(n)/c(nmax)
xx(n)=b(n)-bmin
yy(n)=c(n)-cmin
! xx(n)=b(n)
! yy(n)=c(n)
enddo
valuemin= -emin
valufudge = 1.0d0/(emax-emin)
do k=1,nmax
do j=1,nmax
vertex(j,k)=fepoint(j,k)+valuemin
vertex(j,k)=vertex(j,k)*valufudge
enddo
enddo
do k=1,nmax-1
do j=1,nmax-1
cell(j,k)=fescalar(j,k)+valuemin
cell(j,k)=cell(j,k)*valufudge
enddo
enddo
call XMLwriter01(nmax,xx,yy,cell,vertex)
stop
END PROGRAM BiPolytrope
Subroutine XMLwriter01(imax,x,y,cell_scalar,point_scalar)
real x(200),y(200),z(1)
real cell_scalar(199,199),point_scalar(200,200)
integer imax
integer extentX,extentY,extentZ
integer ix0,iy0,iz0
integer norm(200,3)
! imax=200
ix0=0
iy0=0
iz0=0
extentX=imax-1
extentY=imax-1
extentZ=0
z(1) = 0.0
! Set normal vector 1D array
do i=1,imax
norm(i,1)=0
norm(i,2)=0
norm(i,3)=1
enddo
201 format('<?xml version="1.0"?>')
202 format('<VTKFile type="RectilinearGrid" version="0.1" byte_order="LittleEndian">')
302 format('</VTKFile>')
203 format(2x,'<RectilinearGrid WholeExtent="',6I4,'">')
303 format(2x,'</RectilinearGrid>')
204 format(4x,'<Piece Extent="',6I4,'">')
304 format(4x,'</Piece>')
205 format(6x,'<CellData Scalars="cell_scalars" Normals="magnify">')
305 format(6x,'</CellData>')
206 format(8x,'<DataArray type="Float32" Name="magnify" NumberOfComponents="3" format="ascii">')
207 format(8x,'<DataArray type="Float32" Name="cell_scalars" format="ascii">')
399 format(8x,'</DataArray>')
208 format(6x,'<PointData Scalars="colorful" Normals="direction">')
308 format(6x,'</PointData>')
209 format(8x,'<DataArray type="Float32" Name="colorful" format="ascii">')
210 format(6x,'<Coordinates>')
310 format(6x,'</Coordinates>')
211 format(8x,'<DataArray type="Float32" format="ascii" RangeMin="0" RangeMax="5">')
212 format(8x,'<DataArray type="Float32" format="ascii">')
213 format(8x,'<DataArray type="Float32" Name="direction" NumberOfComponents="3" format="ascii">')
501 format(10f9.5)
502 format(10f9.5)
503 format(5x,9(1x,3I2))
504 format(10f9.5)
505 format(5x,10(1x,3I2))
!!!!!
!
! Begin writing out XML tags.
!
!!!!!
write(*,201) !<?xml
write(*,202) !VTKFile
write(*,203)ix0,extentX,iy0,extentY,iz0,extentZ ! RectilinearGrid
write(*,204)ix0,extentX,iy0,extentY,iz0,extentZ ! Piece
write(*,205) ! CellData
write(*,207) ! DataArray(cell_scalars)
do j=1,imax-1
write(*,501)(cell_scalar(i,j),i=1,imax-1)
enddo
write(*,399) ! /DataArray
write(*,206) ! DataArray(cell_scalars)
do j=1,imax-1
write(*,503)(norm(i,1),norm(i,2),norm(i,3),i=1,imax-1)
enddo
write(*,399) ! /DataArray
write(*,305) ! /CellData
write(*,208) ! PointData
write(*,209) ! DataArray(points)
write(*,502)((point_scalar(i,j),i=1,imax),j=1,imax)
write(*,399) ! /DataArray
write(*,213) ! DataArray(cell_scalars)
do j=1,imax
write(*,505)(norm(i,1),norm(i,2),norm(i,3),i=1,imax)
enddo
write(*,399) ! /DataArray
write(*,308) ! /PointData
write(*,210) ! Coordinates
write(*,212) ! DataArray(x-direction)
write(*,504)(x(i),i=1,imax)
write(*,399) ! /DataArray
write(*,212) ! DataArray(y-direction)
write(*,504)(y(i),i=1,imax)
write(*,399) ! /DataArray
write(*,212) ! DataArray(z-direction)
write(*,504)z(1)
write(*,399) ! /DataArray
write(*,310) ! /Coordinates
write(*,304) ! /Piece
write(*,303) ! /RectilinearGrid
write(*,302) !/VTKFile
return
end
Nonstandard Examination[edit]
In our introductory remarks, above, we said the warped free-energy surface drapes across a five-dimensional parameter "plane" such that,
|
|
|
|
From a more pragmatic point of view, we should have said that the "five-one" free-energy surface drapes across a five-dimensional parameter "plane" such that,
|
|
|
|
In our initial, standard examination of the structure of this warped free-energy surface, we held three parameters fixed — namely, — and plotted . Now, let's fix and plot . The following plot shows how a portion of the grid maps onto the traditional plane. The numerical labels identify lines of constant — 7 (light green), 9 (pink), and 12 (orange) — and lines of constant — 0.330 (purple), 0.315 (dark green), and 0.305 (white/blue).
See Also[edit]
In October 2023, this very long chapter was subdivided in order to more effectively accommodate edits. Here is a list of the resulting set of shorter chapters:
- Free-Energy Synopsis
- Free-Energy of Truncated Polytropes
- Free-Energy of BiPolytropes

|
|---|
|
Appendices: | VisTrailsEquations | VisTrailsVariables | References | Ramblings | VisTrailsImages | myphys.lsu | ADS | |